There is a diversity of statistical methods used for different purposes in microbiological studies. However, some methods are widely known and applied commonly, while other methods even when are suitable for the purpose of research, but are not usually applied because of insufficient knowledge on their existence and interpretation (Windish et al., 2007). The selection and application of statistical methods is closely associated to opportunities provided by the available statistical software.
In order to understand, which statistical software is preferred by the microbiologists, in the database of American Society for Microbiology (ASM), which contains articles of 13 microbiological journals, we analyzed frequency of the use of different statistical packages in published studies:
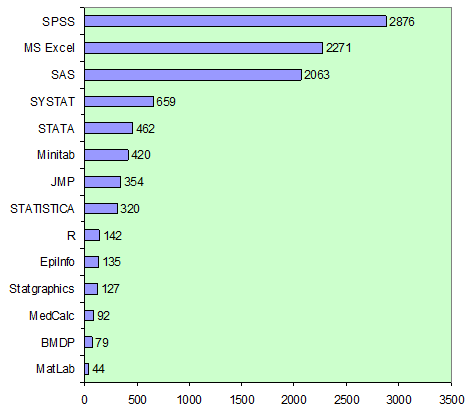
The most frequently used software in microbiological studies is SPSS, although it was originally developed for social sciences. Microsoft Excel is also very common, however, it is almost not applicable for more advanced methods, such as cluster, discriminant analysis etc., and requires additional add-ins and is not recommended by many researchers for statistical analysis of scientific data (McCullough and Heiser, 2008; Yalta, 2008; http://www.practicalstats.com/xlsstats/excelstats.html). The one of the common programs is SAS software and also JMP program, released by the same company.
Most of these software packages belong to commercial. There is also a number of freeware programs, but they are not yet widely used. One of such programs is R software, which has become more popular in the recent years, and in microbiological studies as well (http://www.microbiologybytes.com/blog/statsbytes/).
There are plenty of internet resources with free statistical software, for example http://en.freestatistics.info/stat.php, http://statpages.org/javasta2.html), http://www.statsci.org/free.html, and many other which contain from tens to hundreds statistical programs, general or specialised for some type of analysis:
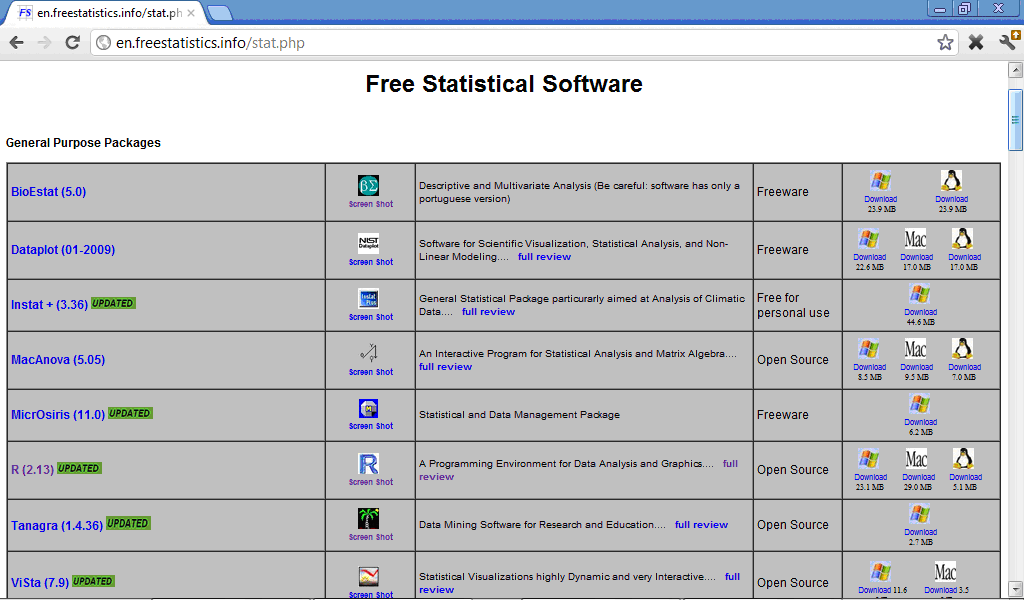
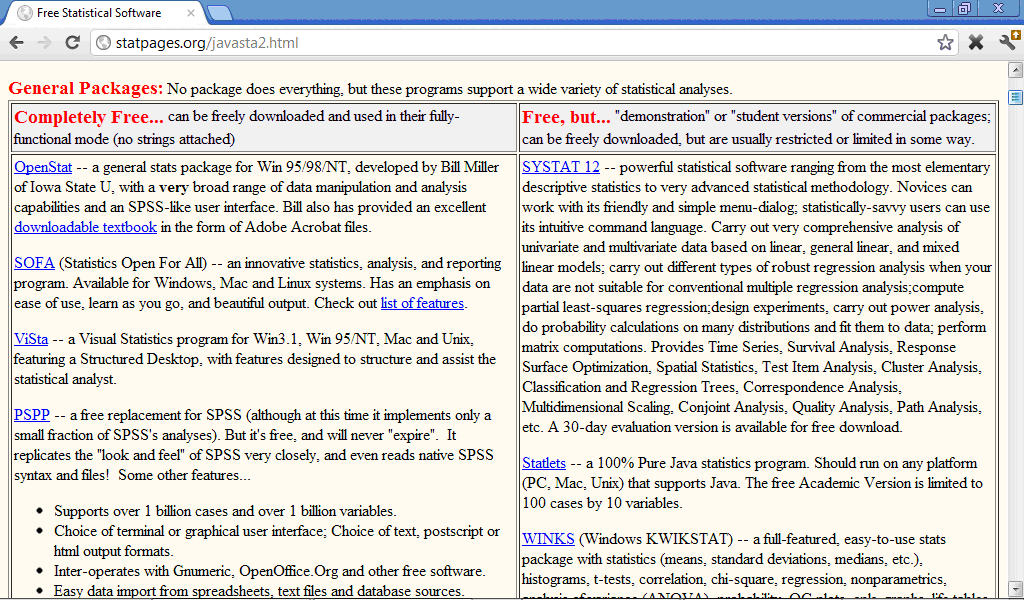
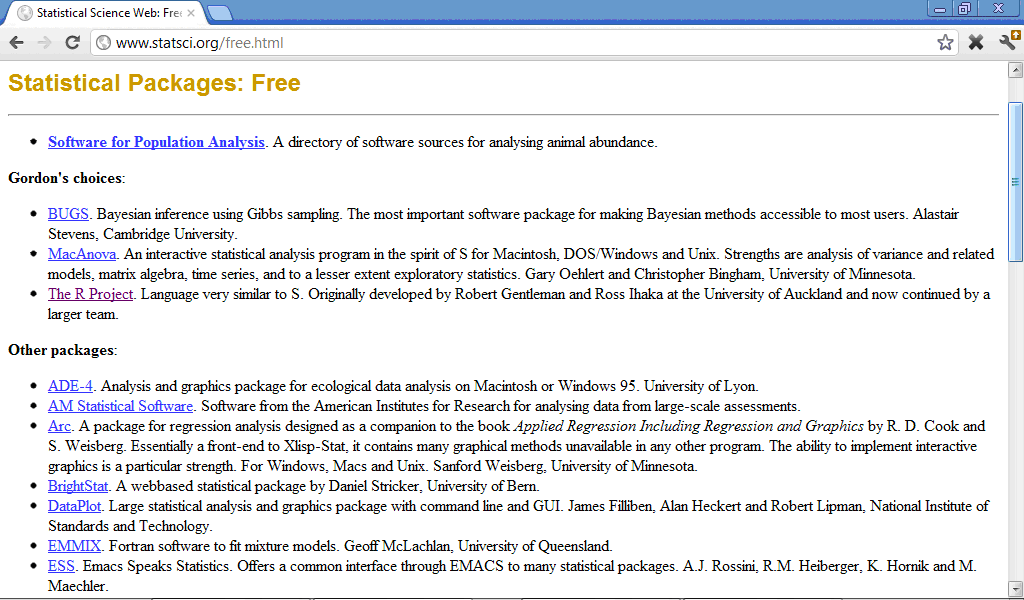
Many of free statistical progrmas, including also R software, require good programming skills because specification of analysis in them must be done through programming language commands and not by clicking correspondent buttons like in SPSS.
Some specialised software packages are very convenient when a researcher has interest in performing particular type of analysis characteristic of biomedical sciences, such as cluster analysis, ROC-curves, etc. For example, program Hierarchical Clustering Explorer (http://www.cs.umd.edu/hcil/hce/) was designed specially for cluster analysis and provides some features absent in SPSS, such as for example interactive comparison of different cluster methods:
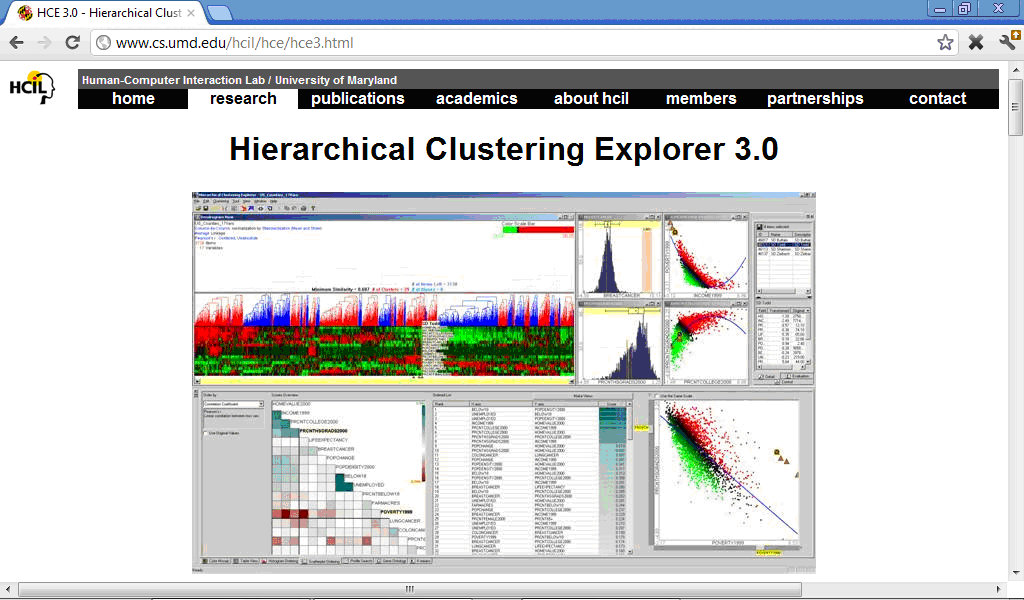
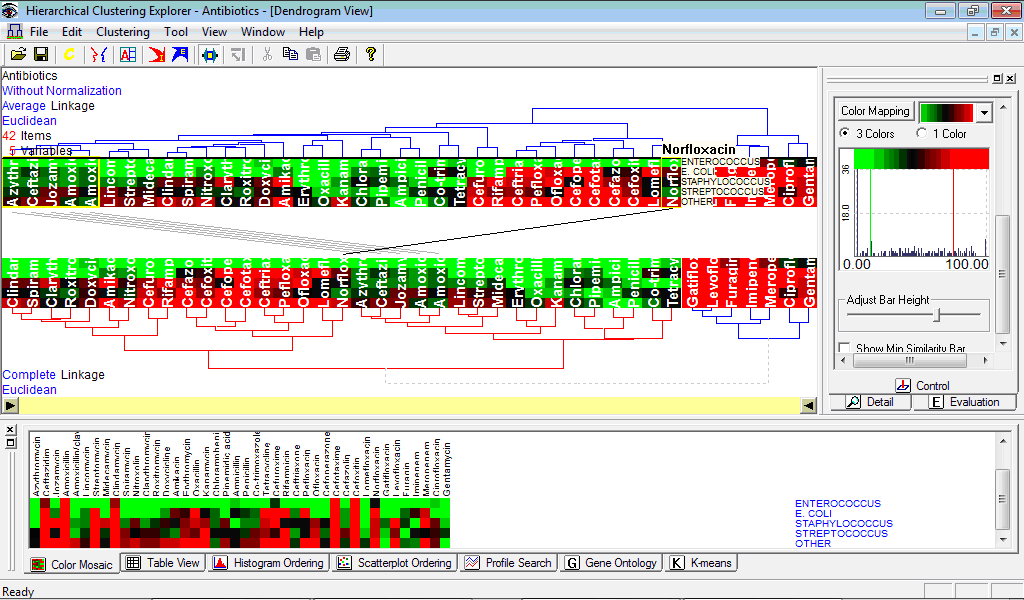
Figure below contains two dendrograms produced for the example data on antibiotic inhibition zones (see dataset here) using average and complete linkages; position of Norfloxacin is interactively located in both dendrograms:
Cluster analysis is mainly used in phylogeny researches, and there are hundreds of specialized programs in this field. One of good internet resources which gathered many available programs is http://evolution.gs.washington.edu/phylip/software.html; it contains links to 388 phylogeny packages and 54 free web servers:
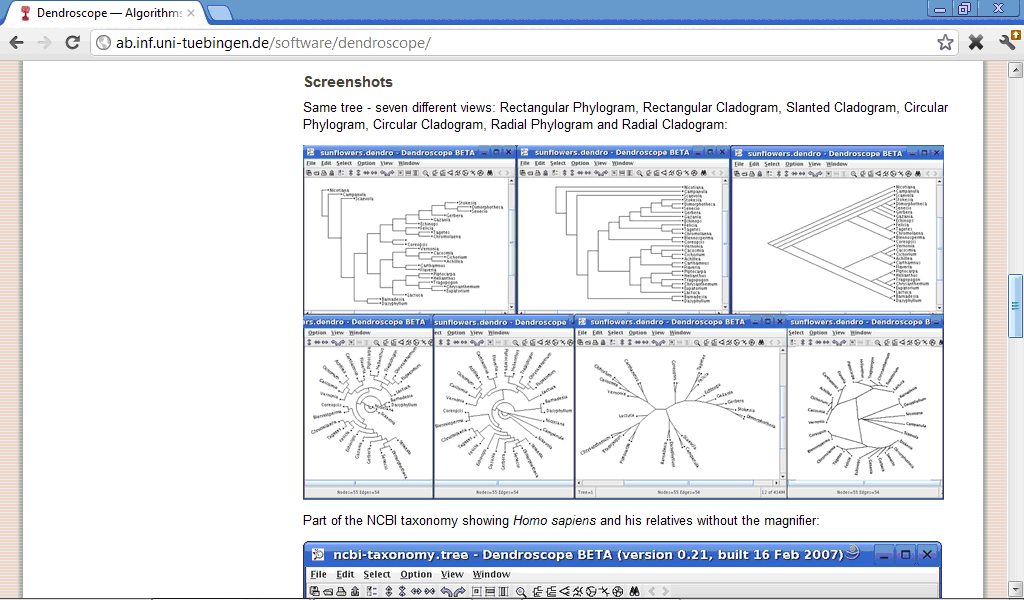
These phylogeny programs provide much more options for performing clustering and building dendrograms, than non-specialised statistical software. For example, program Dendroscope (http://ab.inf.uni-tuebingen.de/software/dendroscope/) provides seven different views of dendrogram, colour highliting of different parts of dendrogram and other features:
There is also free software for analysis of ROC curves, but its amount is not such impressive as for cluster analysis. These are some examples of such programs:
-
Specialized programs for ROC curves: MedRoc (http://www.stenstat.com/), a set of ROC-analysis software developed at the University of Chicago (http://metz-roc.uchicago.edu/MetzROC).
-
ROC curves as part of general statistical software: Tanagra (http://eric.univ-lyon2.fr/~ricco/tanagra/en/tanagra.html), Weka (http://www.cs.waikato.ac.nz/ml/weka/) and Orange (http://orange.biolab.si/) statistical software.
-
On-line web-based calculator for ROC analysis (http://www.rad.jhmi.edu/jeng/javarad/roc/JROCFITi.html).
In order to understand, which statistical methods require greater attention, we analyzed frequency of appearance of different statistical methods in microbiological studies using the reference database of the ASM:
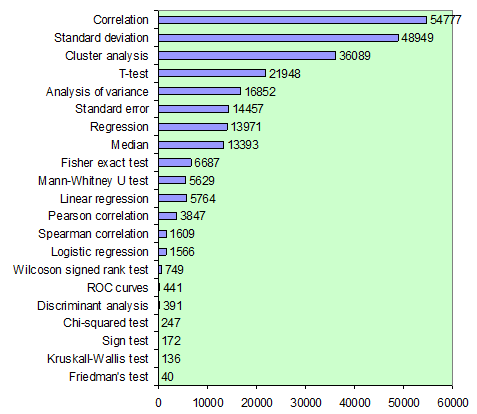
The most commonly used methods were different correlations and descriptive statistics, among which standard deviation is used much more commonly than error of the mean. Many researchers used cluster analysis. For group comparison the most popular is Student’s t-test; however, it must be used only in parametric data, while most biomedical data are non-parametric. Regressions were used almost 4 times rarely than correlations. Logistic regression and ROC curves may provide useful information for researcher but are used very rarely. The least popular methods appeared to be non-parametric analogues of ANOVA – Kruskall-Wallis one-way ANOVA and Friedman’s two-way ANOVA. These figures illustrate the necessity to pay more attention to methods which are still not popular but have high potential in improving quality of microbiological research.