Linear regression model is described by equation
y = a + bx,
where x – value of the first variable, y – value of the second variable, a – intercept of the best fitting line (its position about or below ‘0’ point), b – slope of the line.
Building the best fitting line on a scatterplot is not an easy question because statistical data usually have random scatter and do not lie on a straight line. Random scatter around the line of the best fit is called residuals (vertical lines from dots to the fitting line on the figure below). The line of the best fit is calculated from the goal to minimize the sum of squared residuals and is called the least squares line.
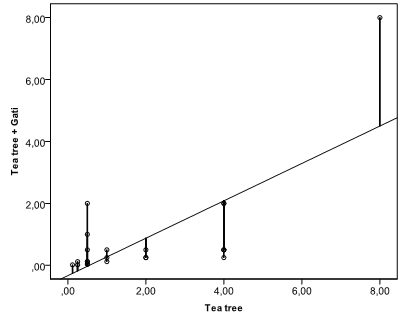
Performing regression analysis
Let us discuss the following example. Optical density of microtubes with Staphylococcus aures, cultivated in the media with different pH and different concentration of glucose (%), was measured in order to evaluate influence of pH and glucose concentration on the growth of S. aureus. The table with dataset includes three columns with values of optical density (OD), pH and glucose concentration:
Optical density of Staphylococcus aureus cultures grown at different pH and different concentration of glucose
|
№
|
OD
|
pH
|
Glucose
|
|
1
|
0.057
|
3.61
|
0.30
|
|
2
|
0.330
|
3.90
|
0.31
|
|
3
|
0.453
|
4.40
|
0.46
|
|
4
|
0.457
|
4.46
|
0.47
|
|
5
|
0.523
|
4.64
|
0.48
|
|
6
|
0.526
|
4.65
|
0.50
|
|
7
|
0.526
|
4.78
|
0.51
|
|
8
|
0.549
|
4.84
|
0.53
|
|
9
|
0.573
|
4.90
|
0.54
|
|
10
|
0.626
|
5.07
|
0.58
|
|
11
|
0.690
|
5.15
|
0.66
|
|
12
|
0.735
|
5.33
|
0.66
|
|
13
|
0.735
|
5.35
|
0.70
|
|
14
|
0.776
|
5.86
|
0.70
|
|
15
|
0.805
|
5.99
|
0.87
|
|
16
|
0.812
|
6.04
|
0.87
|
|
17
|
0.885
|
6.43
|
0.89
|
|
18
|
0.920
|
7.11
|
0.93
|
|
19
|
0.940
|
7.15
|
0.96
|
|
20
|
0.947
|
7.39
|
0.99
|
The research question is: How does growth of S. aureus depend on pH and glucose concentration in the medium? If we study only dependence of optical density on pH, it is simple regression; but when we study dependence on several factors (pH and glucose) – it is multiple regression. In SPSS specifying both simple and multiple regression is similar.
To specify regression analysis:
1. Click the Analyze menu, point to Regression, and select Linear… :
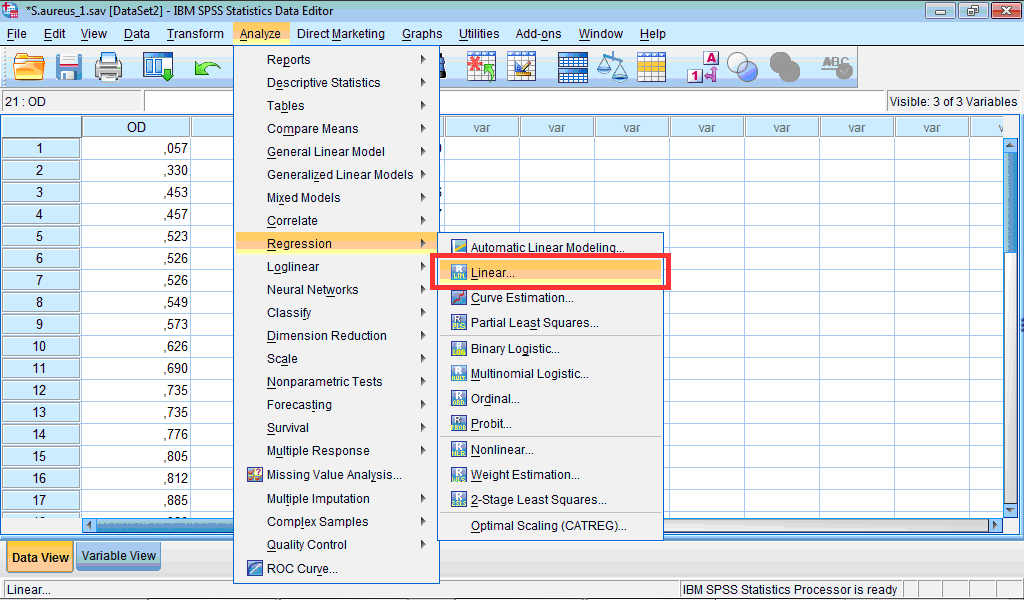
The Linear Regression dialog box opens:
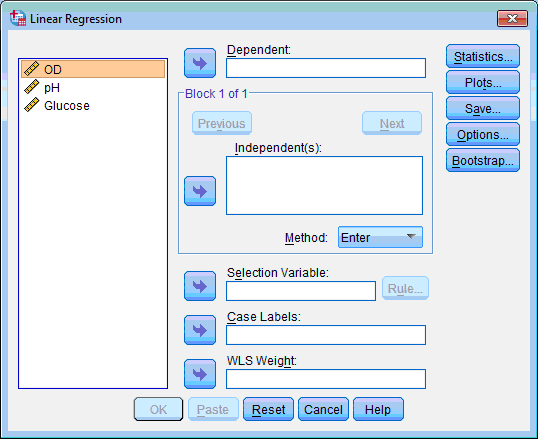
2. Select dependent variable (which we want to predict – “OD”); click the upper transfer arrow button  , the variable is moved to the Dependent: list box.
, the variable is moved to the Dependent: list box.
3. Select independent variable/s (which is/are used for prediction – “pH” and “Glucose”); click the transfer arrow button  from the section Block 1 of 1, the variables are moved to the Independent(s): list box.
from the section Block 1 of 1, the variables are moved to the Independent(s): list box.
4. Click the Statistics… button. The Linear Regression: Statistics dialog box opens:
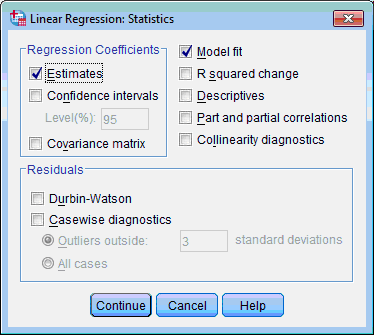
5. By default two check boxes are selected – Estimates and Model Fit. Select also the Part and Partial correlations and Collinearity diagnostics check boxes. Click the Continue button. This returns you to the Linear Regression dialog box.
6. Click the OK button. An Output Viewer window opens and displays results of regression analysis.
In the Output Viewer window five tables appear with results of regression analysis – Variables Entered/Removed, Model Summary, ANOVA, Coefficients and Collinearity Diagnostics tables The table Variables Entered/Removed is important when we use methods of regression other then Enter; in such case it shows which variables were chosen to be included to the model. Because in the Linear Regression dialog box we did not change selected by default “Enter” method, all specified by us variables were included:
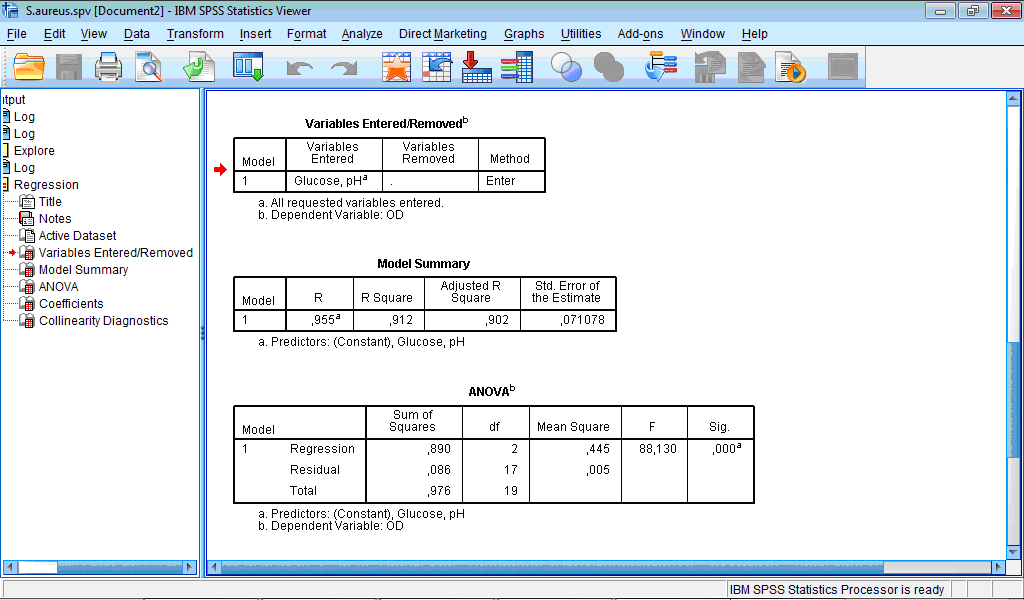
The Model Summary table contains R values. R squared shows that the model works very well and explains more than 90% of the variation in optical density.
The ANOVA table reports a significant F statistics (last two columns), which indicates that the use of the model is better than guessing by chance.
The Coefficients table contains information necessary to build the regression model itself and also to assess the presence of correlations between variables:
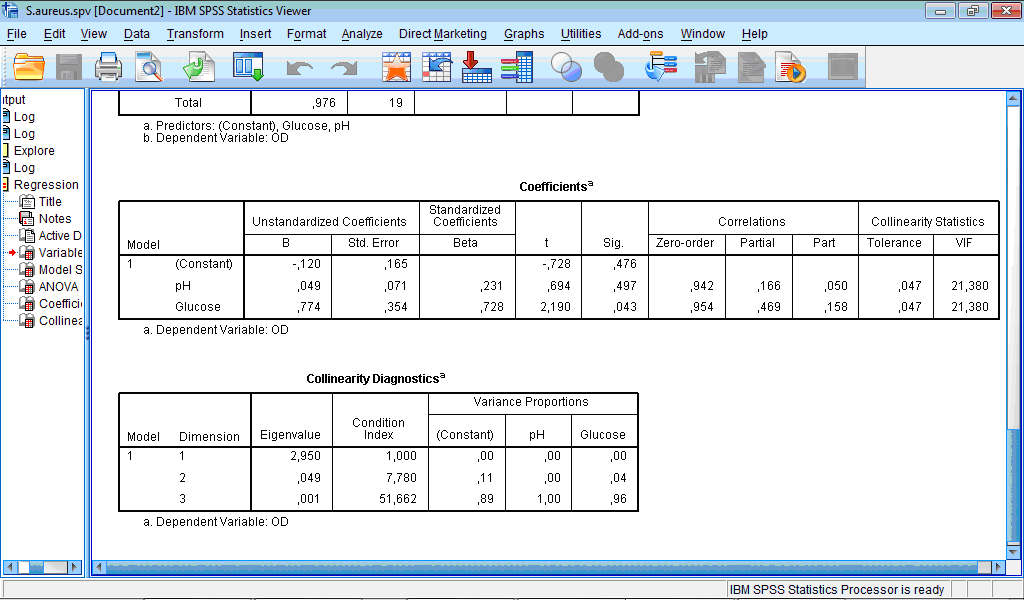
The coefficients for regression model present in the first column (“B”), that is, our model can be written as
OD = -0.12 + 0.049×pH + 0.774×Glucose.
From this equation we may conclude that optical density increases with increase either pH or glucose concentration, i.e., S. aureus better grows at higher pH and glucose content.
However, evaluation of all columns is important to understand the required modifications to the model. Variable/s with non-significant coefficients do not contribute much to the model. In our example it is “pH” variable. It is also seen from the column with standardized coefficients: coefficient for glucose is higher than coefficient for pH – 0.728 and 0.231, respectively.
Columns with correlations and collinearity statistics show that there may be a problem with multicollinearity. When values of partial and part correlations drop sharply from the zero-order correlation, it means that, for example, much of variance in optical density that is explained by pH is also explained by other variables (glucose). Tolerance is the percentage of the variance in a given predictor that cannot be explained by the other predictors. Low values of tolerance (even less than 5%) shows that most of the variance in a given predictor can be explained by the other predictor. Value of tolerance close to 0 indicates high multicollinearity, and in such case the standard error of the regression coefficients will be inflated. “VIF” is a variance inflation factor, usually its value more than 2 is considered as a problematic. In our example VIF values are very high which also indicates high multicollinearity and the necessity for further transformations of variables.
The table Collinearity Diagnostics further proves high multicollinearity of the variables: eigenvalues close to zero indicate that the predictor variables are highly intercorrelated and small changes in the data values may lead to large changes in the estimates of the coefficients. The condition indices (next column) are computed as the square roots of the ratios of the largest eigenvalue to each successive eigenvalue and their high values (more than 15) also indicate serious problem with collinearity between predictors.
Problems with collinearity can be solved in different ways:
1) Values of variables can be transformed to standardized (so called z-scores). Z-score transformation standardizes variables to the same scale, producing new variables with a mean of 0 and a standard deviation of 1.
2) When there are many variables, they can be transformed to fewer new variables by factor analysis.
3) Regression with automatic selection of variables can be used (stepwise, forward or backward).
First, let us try to produce z-scores of predictor variables and to built a new model based on these z-scores.
1. Click the Analyze menu, point toDescriptive Statistics and select Descriptives… The Descriptives dialog box opens:
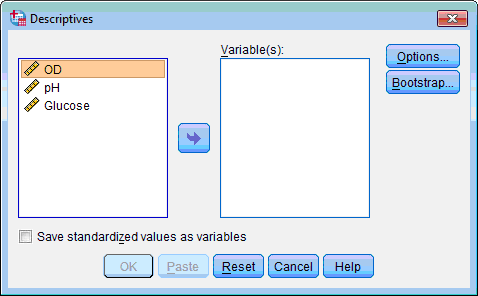
2. Select the variables to be standardized (“pH” and “Glucose”), click the transfer button , the variables are moved to the Variable(s): list box.
3. Select Save standardized variables as values check box.
4. Click the OK button. Two new columns with standardized variables appear in the Data Editor window:
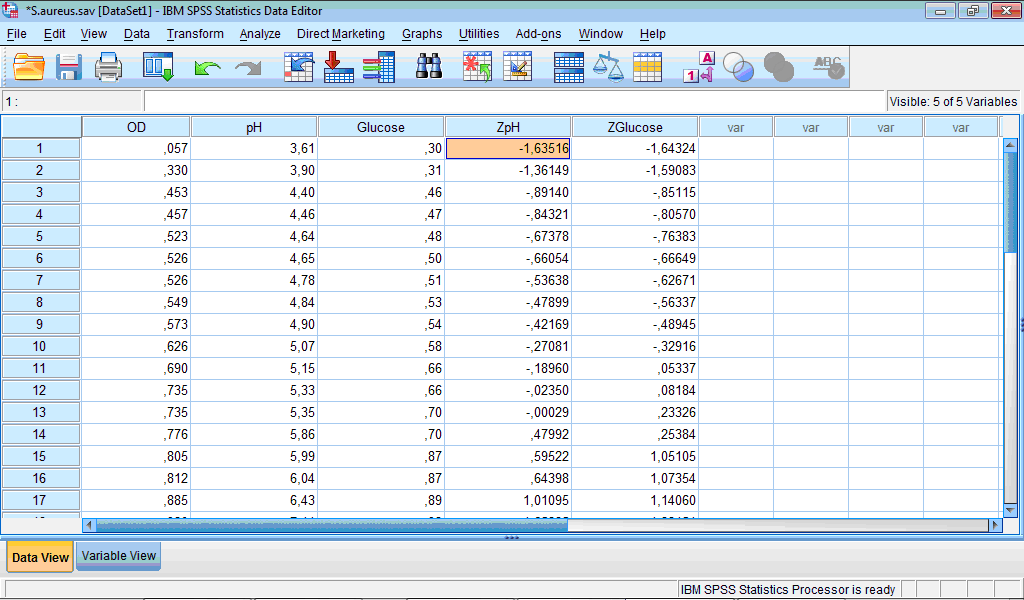
Linear regression analysis is specified in the same way as described above, but for the independent predictors z-scores are selected:
Compared to original model, eigenvalues and condition indices are vastly improved:
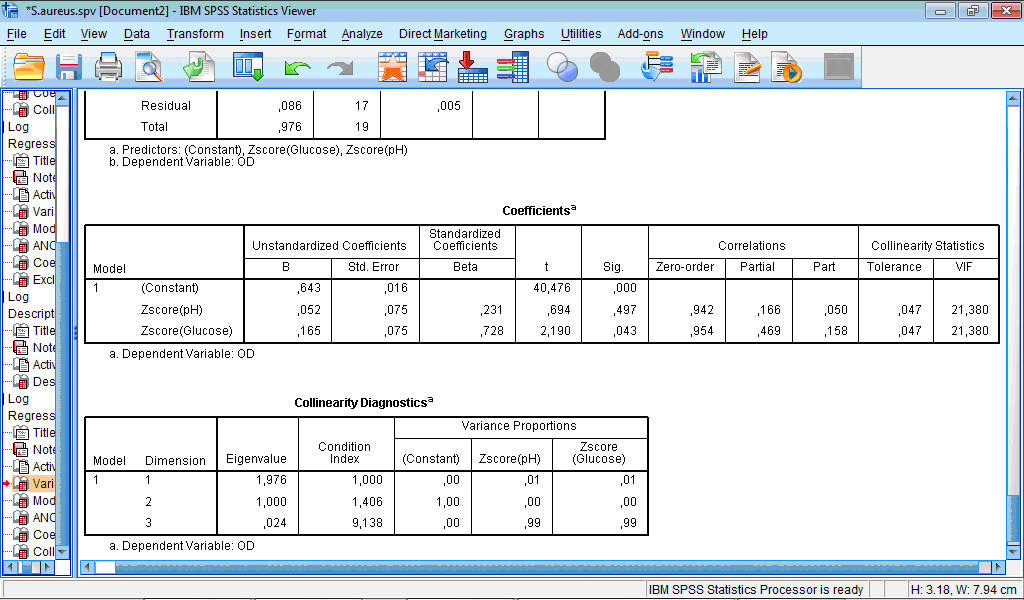
However, correlations and collinearity statistics still indicates presence of problems with multicollinearity. Z-score transformation does not change correlation between variables and because of this collinearity statistics in unimproved. The condition index, which is used in multicollinearity diagnostics, is important for flagging datasets that could cause numerical estimation problems in algorithms but do not internally rescale the independent variables. Such problem can be solved by the z-score transformation. However, this procedure appears not sufficient in our case.
We have only two predictor variables and because of this we can manually include more significant one (“Glucose”) into the model. In general, to include/exclude less significant variables we may use step-wise, forward or backward regression with specification of criteria for including/excluding. Let us also illustrate the use of forward regression for inclusion of the most statistically significant variables. It is worth to mention, that the choice of variables is performed by statistical significance but there may be no practical significance. Because of this, after performing analysis included and non-included variables should be carefully examined for making the sense.
During specification of regression analysis, in the Method drop-down menu select the Forward method:
By clicking the button Options we may specify including criteria or may use proposed by default F value of 0.05:
In Linear Regression: Statistics dialog box deselect Part and Partial correlations and Collinearity diagnostics check boxes.
The Output Viewer window contains the following tables: Variables Entered/Removed, Model Summary, ANOVA, Coefficients and Excluded Variables:
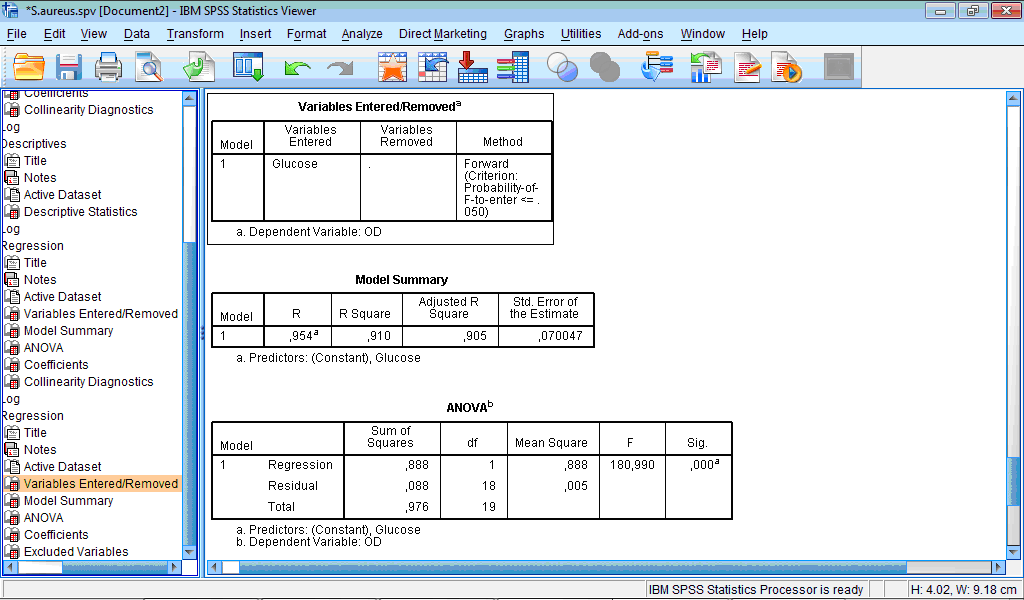
All except last table are built by the same principle like for above described regression.
From the table Variables Entered/Removed we can see that variable “Glucose” was entered and the model is built based on this variable.
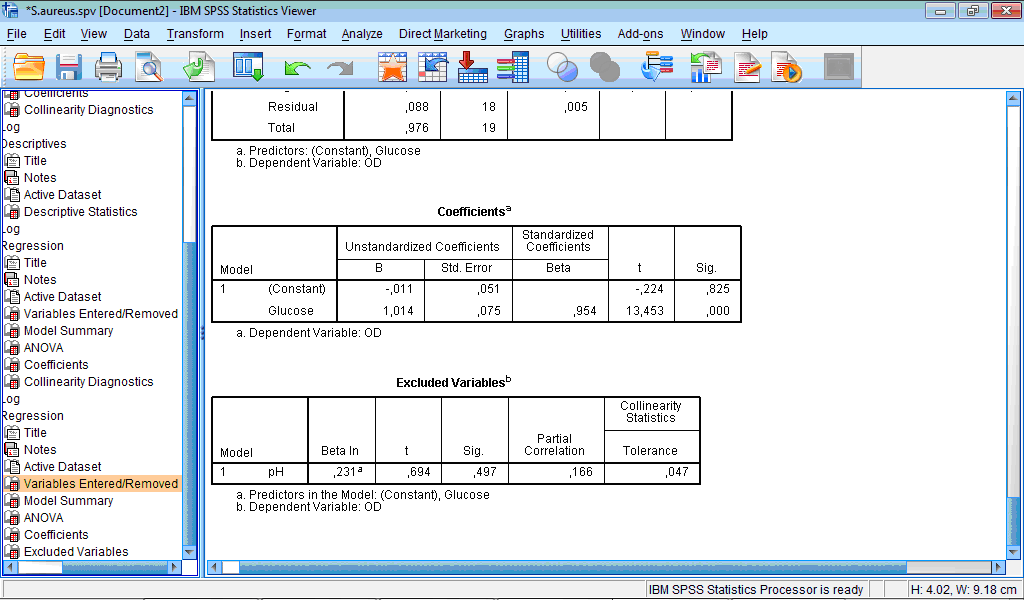
The regression model is: OD = -0.011 + 1.014 * Glucose. This is seen from the Coefficients table.
Therefore, OD is almost equal to the glucose content in the medium. From microbiological point of view it may be explained by stimulation of the growth of S. aureus, which corresponds to the increase of optical density of culture solution, caused by increasing concentration of glucose.